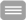Research activities
Main topic: Karyotype evolution
We focus on the following aspects of karyotype evolution:
- Comparative whole genome karyotype analysis of great apes (chimpanzee, gorilla and orang utan) by high resolution mMCB and subCTM FISH.
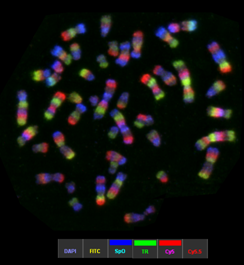
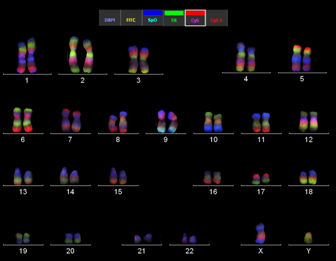
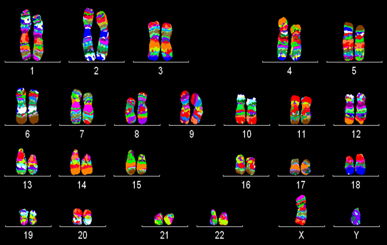
Example of a mMCB hybridization on a normal male metaphase spread. From left to right: metaphase spread, the corresponding karyogram and pseudocolor image. This high resolution technique is currently used in Zoo-FISH studies on great apes to uncover cryptic until now overlooked rearrangements and furthermore to narrow breakpoint regions for subsequent breakpoint mapping by BACs.
Application of mMCB to gorilla metaphase spreads revealed the the same results as with single MCB hybridizations (Mrasek et al., 2001). As described in Weise et al., 2003 the analysis of mMCB is primary done on fluorescence profiles and than followed by pseudocolor evaluation.
Further topics
-
mapping and characterization of evolutionary fixed breakpoints by BACs from the human genome project.
-
mapping and characterization of fragile sites and comparison to cancer and evolutionary breakpoints.
-
glass needle microdissection based investigation of different kind of heterochromatin in great apes.
-
mapping and characterization of human inversion variants.
-
karyotype evolution in different cell lines from adenoma to tumour.
-
M-FISH, mMCB and MCB based cell line characterization
Motivated and interested biology and medical students are encouraged to contact Dr. Anja Weise for diploma or PhD thesis projects
Publikation Link